Downloading Traces from LIMS
The Geneious Assembler Module is used to edit, save, export and ultimately publish the sequence data. To be able to use the Assembler Module, traces must be imported into the user’s local Geneious Prime directory.
There are two options for importing the raw traces:
Downloading the raw traces from the LIMS database
Directly importing the traces from local disk
The main advantage of the first option is that all of the specimen’s associated FIMS and LIMS metadata are attached to its trace file. Alternatively, if importing traces from disk, none of the specimen’s associated FIMS or LIMS metadata will be attached, and it can prove more difficult to attach this metadata after import. As a result, the first option should be utilized whenever possible.
Downloading the Raw Traces from the LIMS Database
Log in to the Biodcode LIMS Plugin. See Connecting with the Biocode Plugin for instructions.
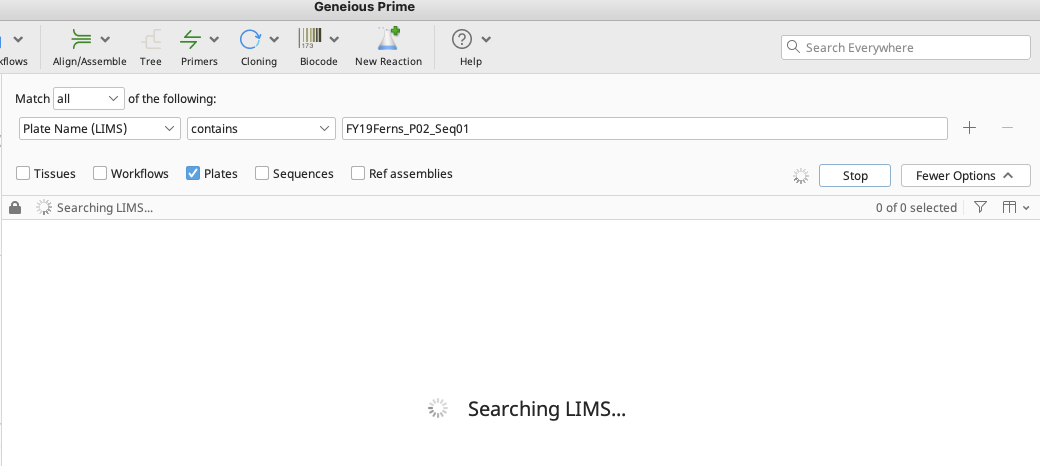
Search for the relevant cycle sequencing plate(s) in the Biocode LIMS.
Select the relevant plate(s) from the search results.
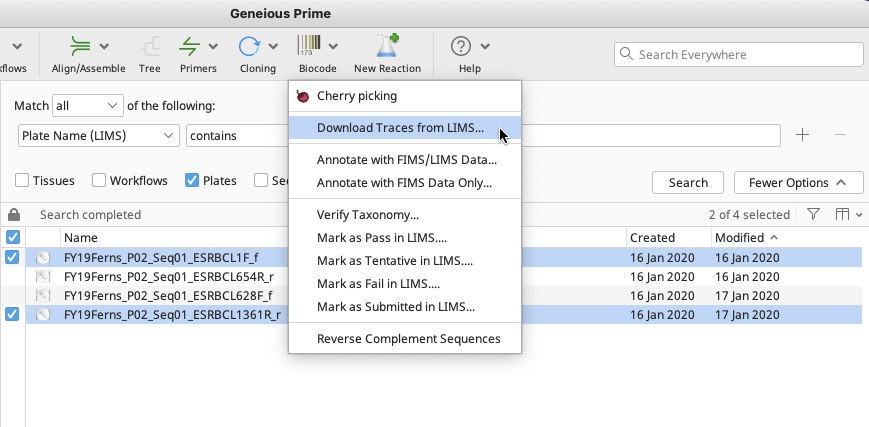
Click the “Biocode” button in the Toolbar and choose “Download Traces from LIMS…” to begin the operation. A Select Destination window will open.
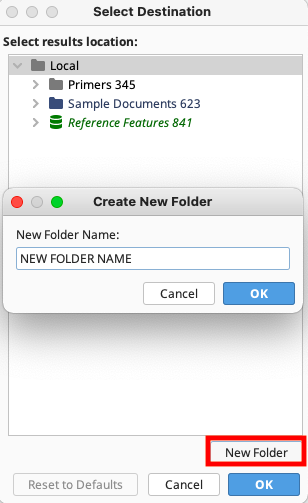
Choose a destination folder in the local directory for the downloaded traces. Either select an existing folder or create a new folder. To generate a new folder, highlight the folder in the local directory in which the new folder will be located, select “New Folder”, and provide a name.
Once the destination folder has been selected, click “OK” and the Download Traces from LIMS window will pop up with the names of the previously selected plates automatically filled in.
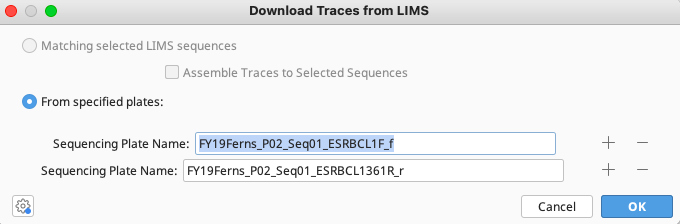
Click “OK” on this window to begin downloading the trace files into the designated folder.
Alternatively, if the exact names of cycle sequencing plates are known, it is possible to download the plates directly without having to perform a search. To do this:
Log in to the Biodcode LIMS Plugin.
Highlight a destination folder under the Local Directory.
Select the “Biocode” button in the Toolbar followed by the “Download Traces from LIMS…” option and enter the cycle sequencing plate name manually.
If downloading traces from more than one plate, use the “+” button to add more fields for the additional plate names.
Once this is complete, click “OK “and Geneious will begin downloading the trace files into this folder.
Directly Importing the Traces from Disk
There are two ways to do this, both must be followed up by the third section below:
Using the Geneious Prime Toolbar
In the Geneious Prime local directory, create a destination folder for the raw traces, and highlight it.
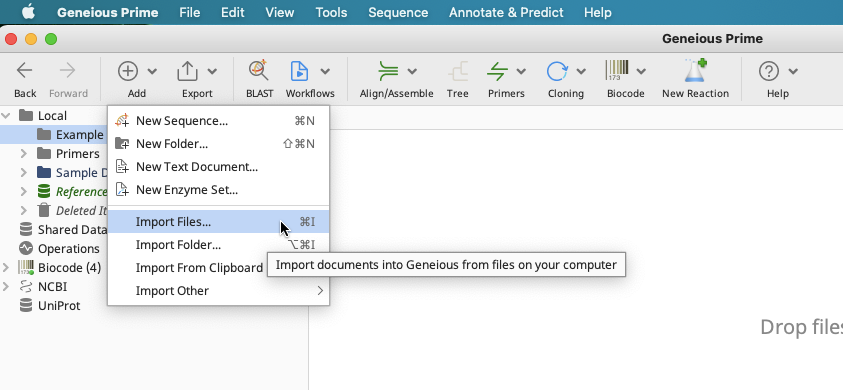
In the Toolbar, click on “Add” > “Import Files…”.
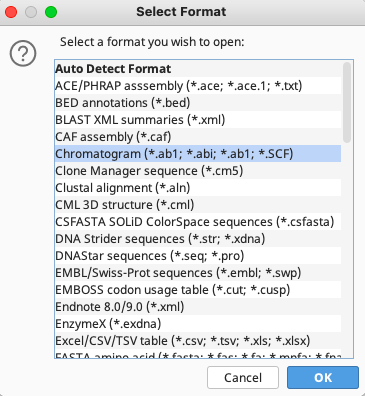
In the Select Format window that appears, select “Chromatogram(*.ab1; *.abi; *.ab1; *.SCF)” then click “OK”.
Choose the directory on the local disk where raw traces are located.
Click “Open” and the trace files will be imported.
Drag-and-Drop
Outside of Geneious Prime, navigate in the computer’s file explorer to the trace files.
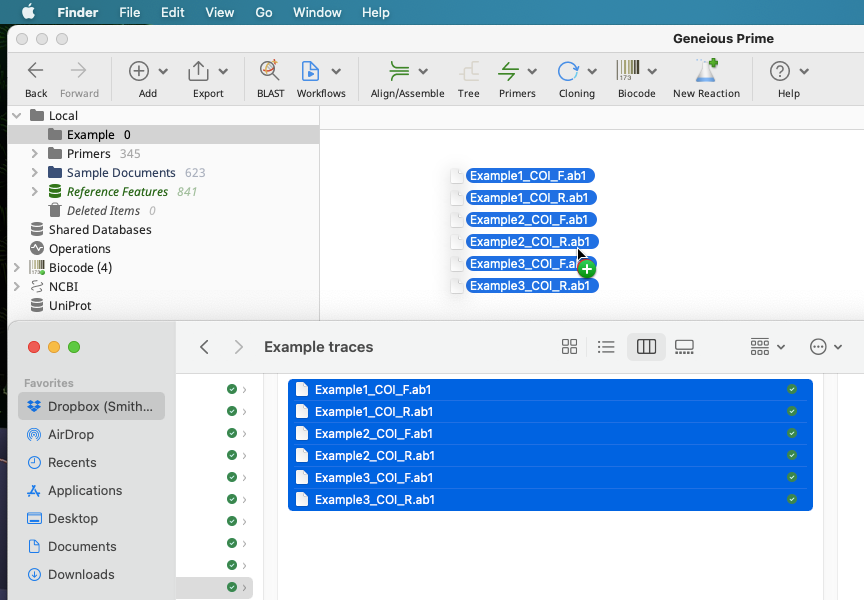
Select the trace files to import, and drag them into a selected folder in the local directory of Geneious Prime.
Read Direction and Metadata
Once the raw trace files have been imported from the local disk, it is necessary to define the read direction. To do this:
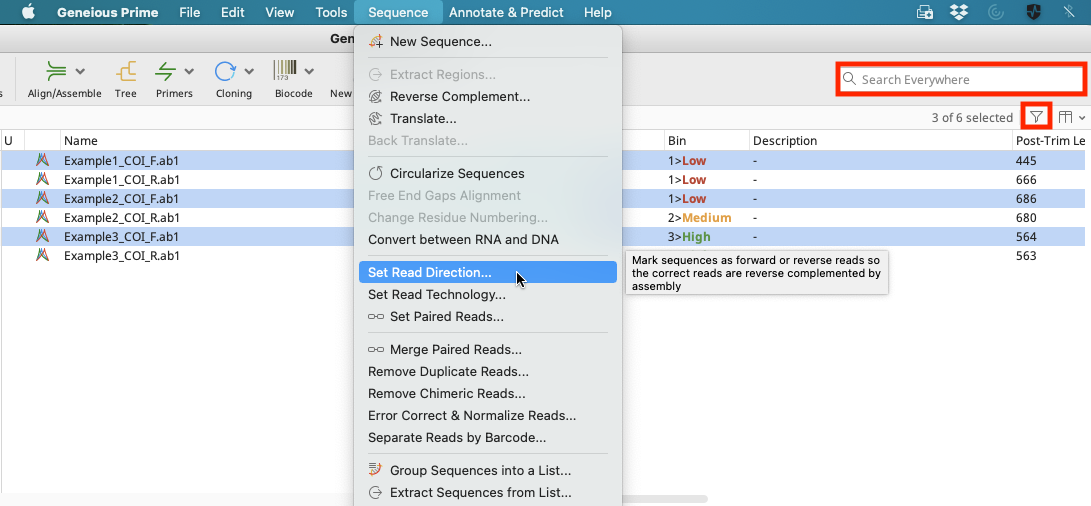
Choose either the forward or reverse trace files and select “Sequence” in the Menu bar followed by “Set Read Direction”.
Choose the read direction and click “OK”. It is only necessary to choose the direction for one set of reads because the other set of reads will be assigned the opposite direction by default.
After performing this task, an extra column will be added to the Document Table titled, “Is Forward Read” with a value of true or false.
If the forward and reverse traces are in different folders, it’s easiest to import all of the traces from one folder, set the read direction for that folder and then import the second folder.
The Search dialog box or Filter button in the upper right-hand corner of the Geneious Prime window can be used to locate a particular direction of reads based on trace names if both forward and reverse traces are imported in one folder. Both options are highlighted in the above screen shot.
Traces imported into Geneious directly from disk have none of the specimen’s associated FIMS or LIMS metadata attached. It is possible to annotate traces with the associated metadata, but this must be done pre-assembly (with the traces) because forward and reverse traces can come from different sequencing plates.
Note
It is more difficult to use the “Annotate from FIMS/LIMS Data” function on traces where both forward and reverse come from the same cycle sequencing plate. In these cases, it is strongly recommended that the user only download traces from the LIMS and not upload directly from a local disk.
To attach the associated metadata:
Click the “Biocode” button in the Toolbar, then select “Annotate from FIMS/LIMS Data…”.
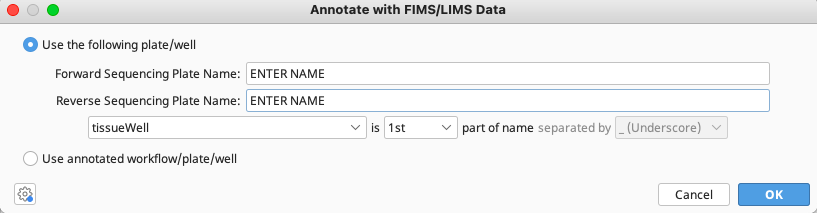
A new window appears to enter the forward and reverse sequencing plate names (from the LIMS) that correspond to the traces. The location of the FIMS field tissueWell and delimiter in the trace name must also be indicated.
Click “OK” and the operation will run.
For further instruction on annotation other local document types with FIMS or LIMS metadata, see the Annotating with FIMS/LIMS Data page.