NCBI BioProjects
What is a BioProject?
From the NCBI BioProject homepage (https://www.ncbi.nlm.nih.gov/bioproject/):
“A BioProject is a collection of biological data related to a single initiative, originating from a single organization or from a consortium. A BioProject record provides users a single place to find links to the diverse data types generated for that project.”
BioProjects grew out of the NCBI Genome Project database, which served solely to organize genome sequences in GenBank. However, it became apparent that this organizational structure could be used to group together entries for several more kinds of data, so BioProjects became a distinct entity in 2011.
BioProjects can have a hierarchical structure, meaning that project-level BioProjects can be organized together under an “umbrella” BioProject.
The Smithsonian Barcoding Network (SIBN) uses BioProjects to organize sequence records on GenBank that were generated by each project that it funded. Each funded project will have its own BioProject that makes searching easier and tracking progress more convenient. The SIBN BioProject can be found at https://www.ncbi.nlm.nih.gov/bioproject/81359,
When a GenBank record is added to a BioProject, a link to other records in the same BioProject appears directly on the GenBank record.
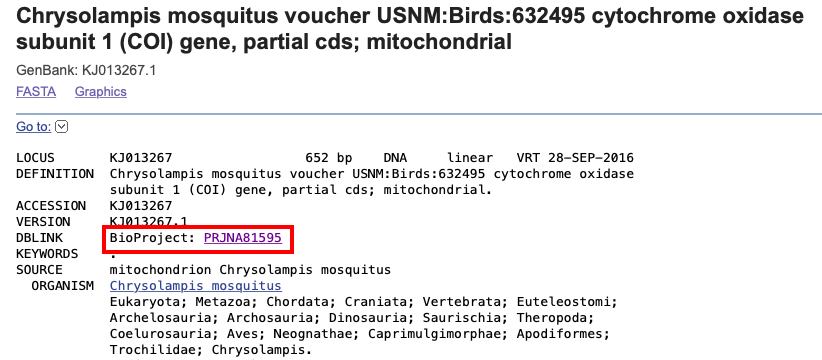
Here is an example of a BioProject link appearing in a GenBank record (https://www.ncbi.nlm.nih.gov/nuccore/KJ013267)
Creating a BioProject
A BioProject will need to be created before new GenBank submissions or existing GenBank records can be organized under one.
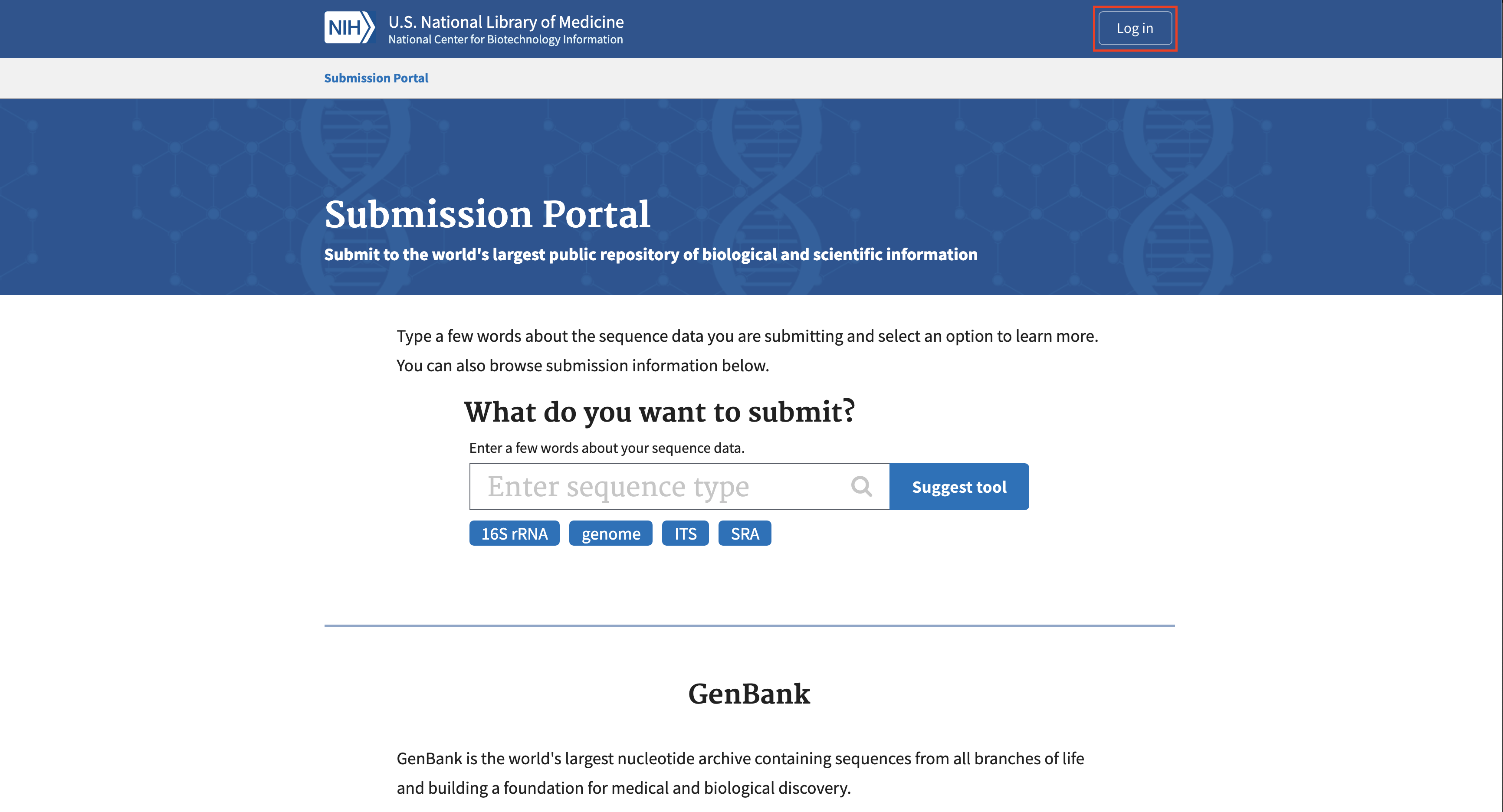
To create a BioProject, navigate to the NCBI Submission Portal: https://submit.ncbi.nlm.nih.gov/.
Click on “Log in”. Generally, it is easiest to sign in with a Google or other third party account. An account can also be made with Smithsonian credentials.
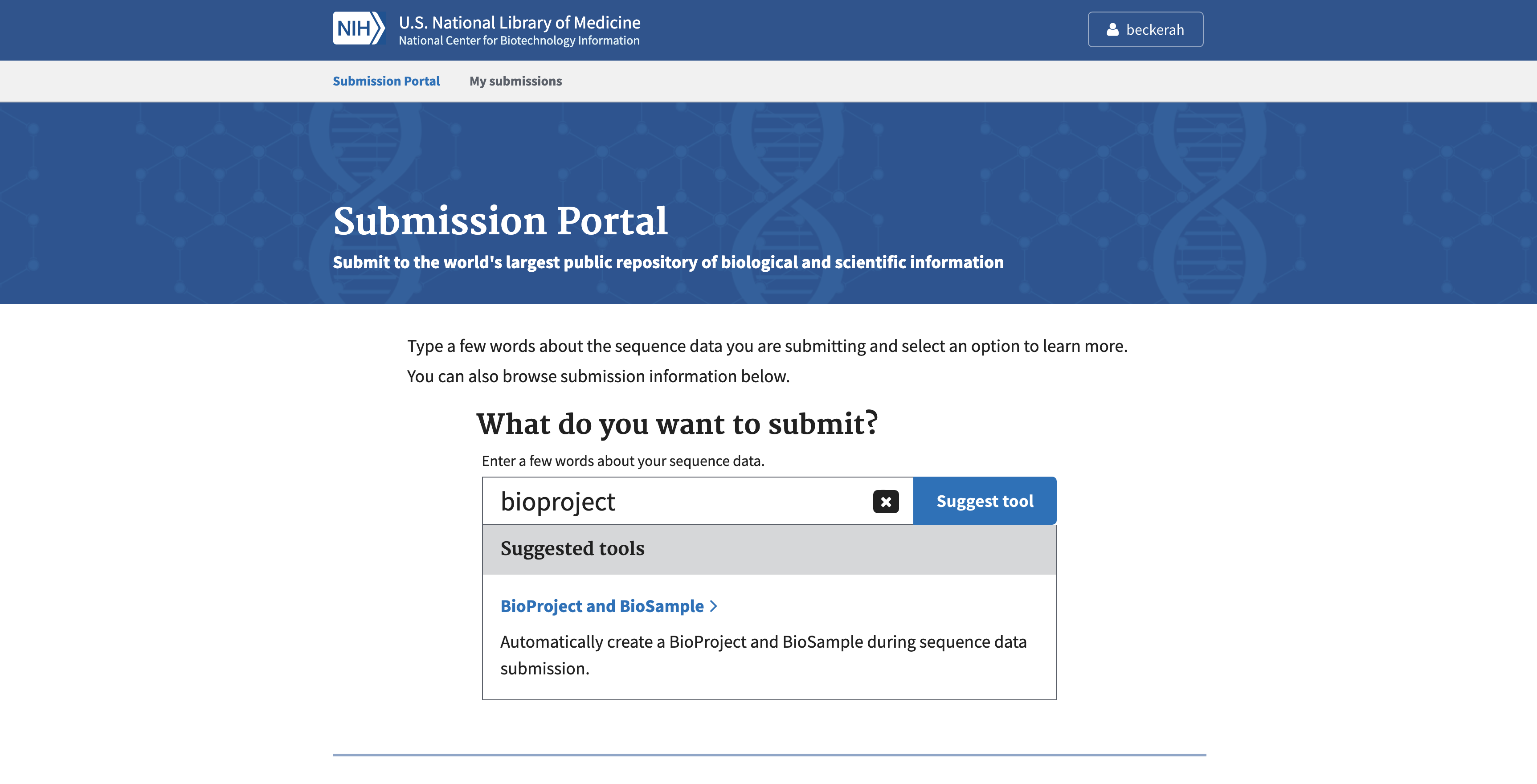
After signing in, return to the Submission Portal page. In the search box, type “bioproject” and click on the link “BioProject and BioSample”. This will lead to the BioProject submission tool.
Feel free to click through the headings on the left of the “What You Should Expect” section to learn more about the requirements and the submission process. When ready to continue, click the Submit button.
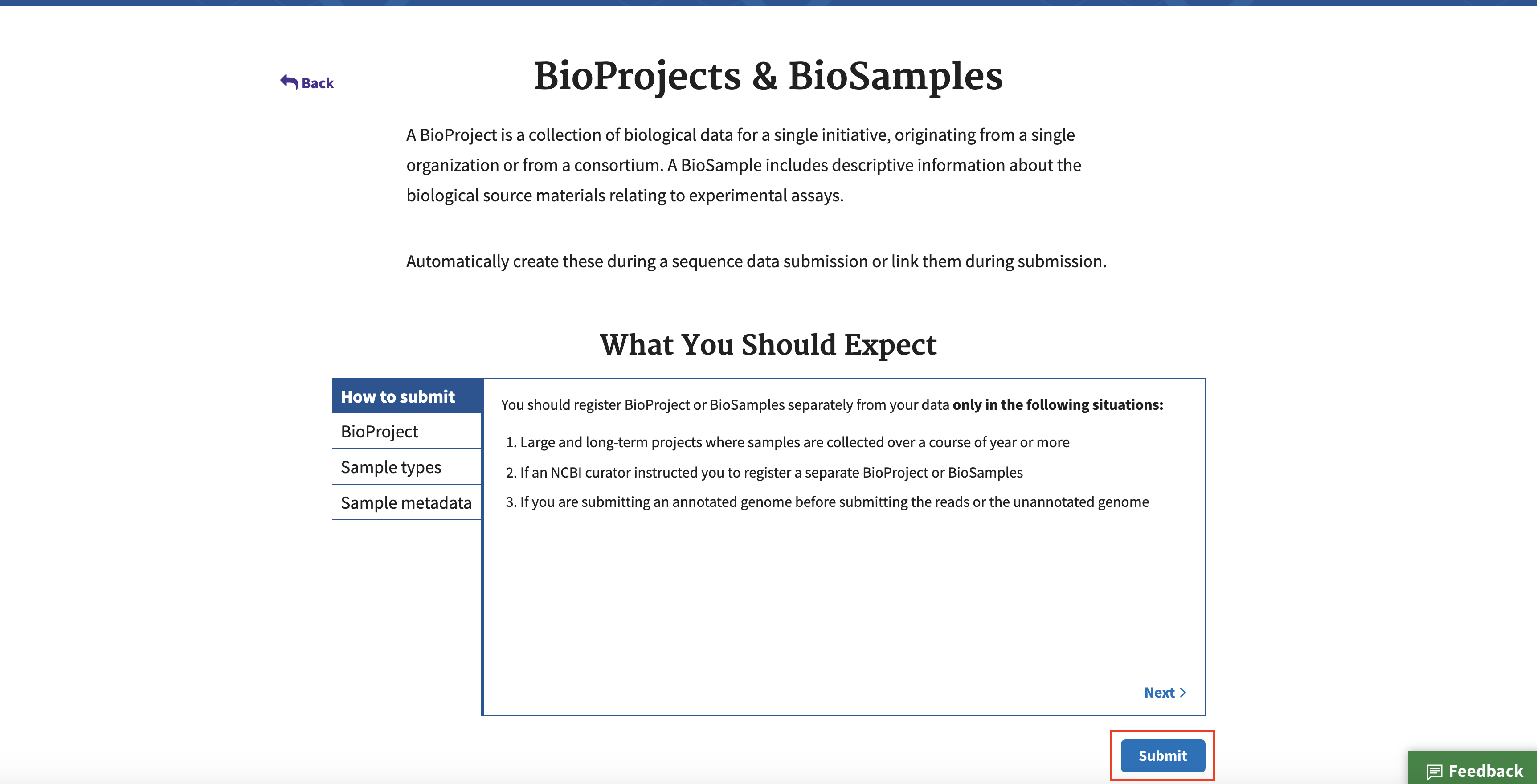
Click the New Submission button. A series of 7 tabs to be filled out will appear.
Submitter Tab
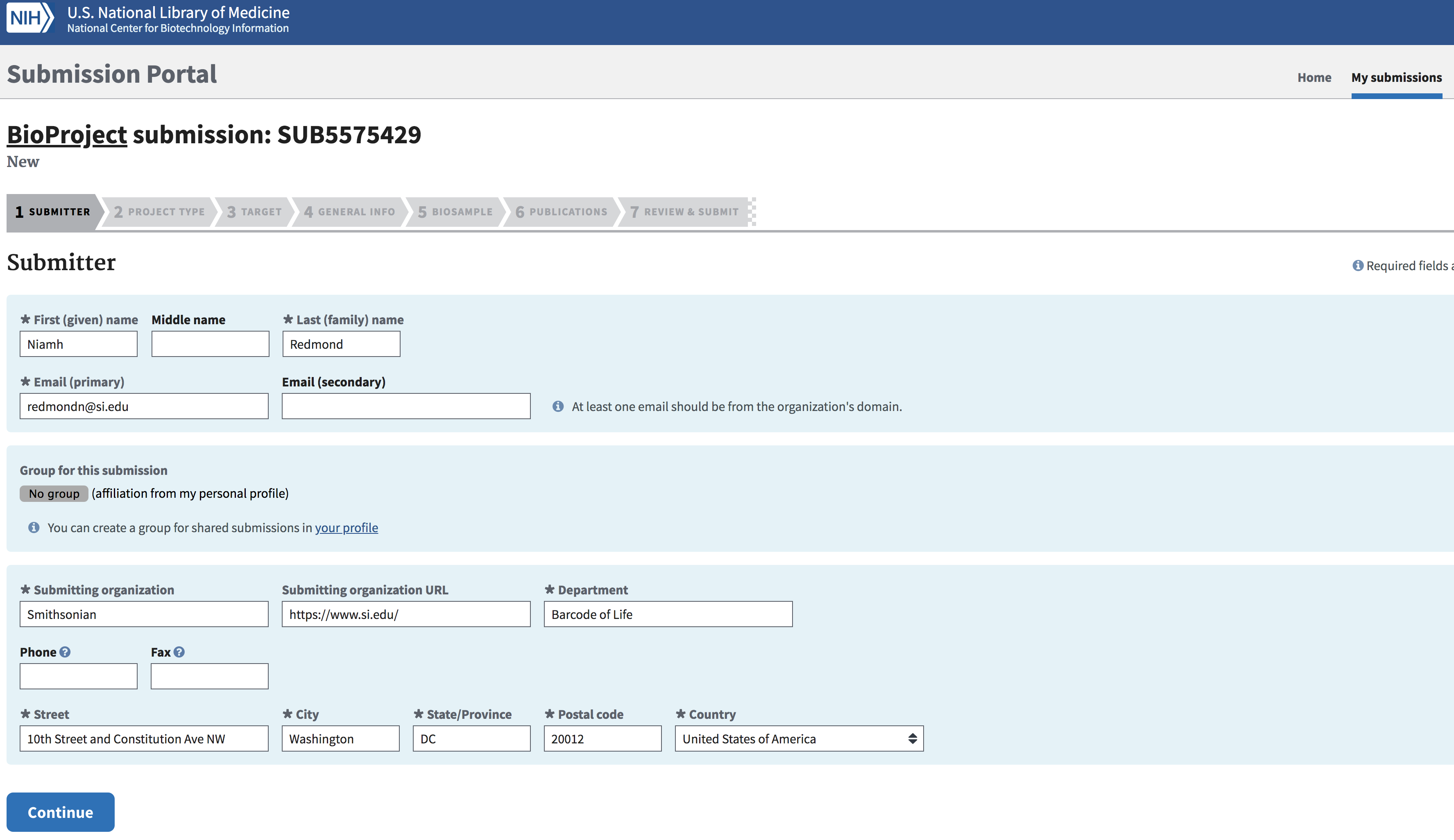
Fill out the Submitter page. Click “Continue” when ready.
Project Type Tab
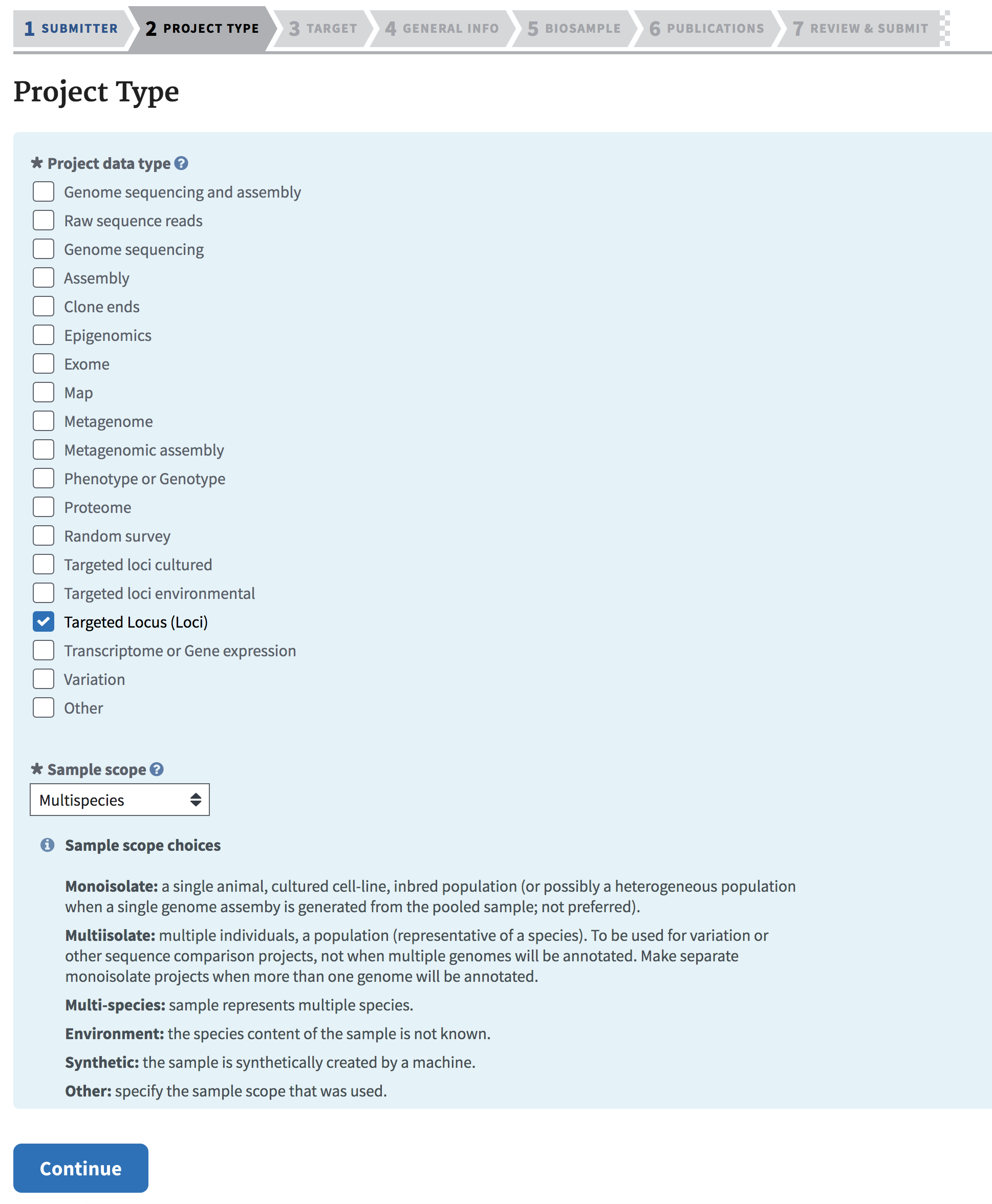
For traditional DNA barcoding projects, select “Targeted Locus (Loci)” for Project Data Type. For projects that will contain assembled genomes and/or raw reads in the SRA, select “Genome sequencing and assembly” and/or “Raw sequence reads”.
Typically, SIBN funded projects involve sequencing the same markers across many taxonomically different samples, so select “Multispecies” for Sample scope. Click “Continue” when ready.
Target Tab
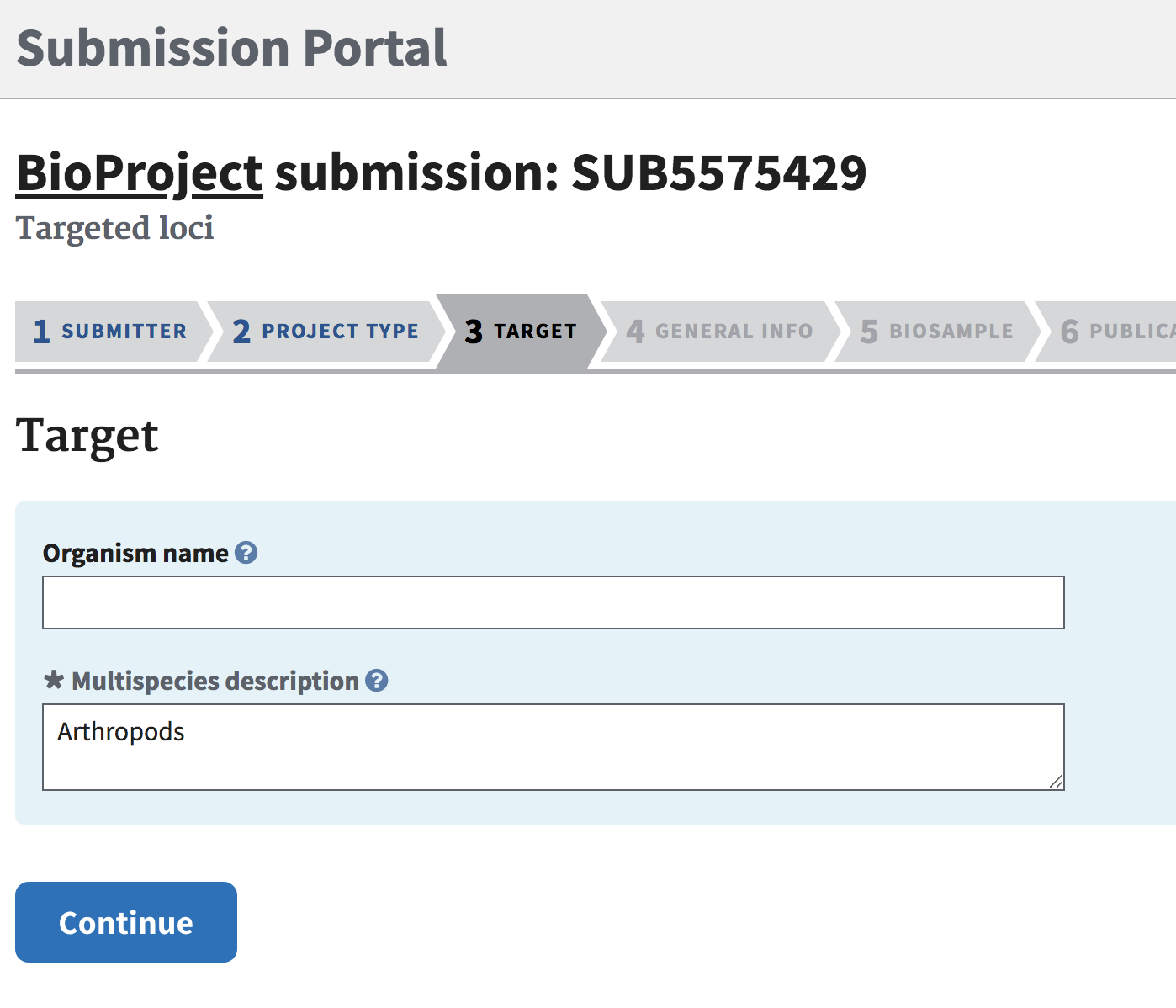
Give a short description for “Multispecies description”.
General Info Tab
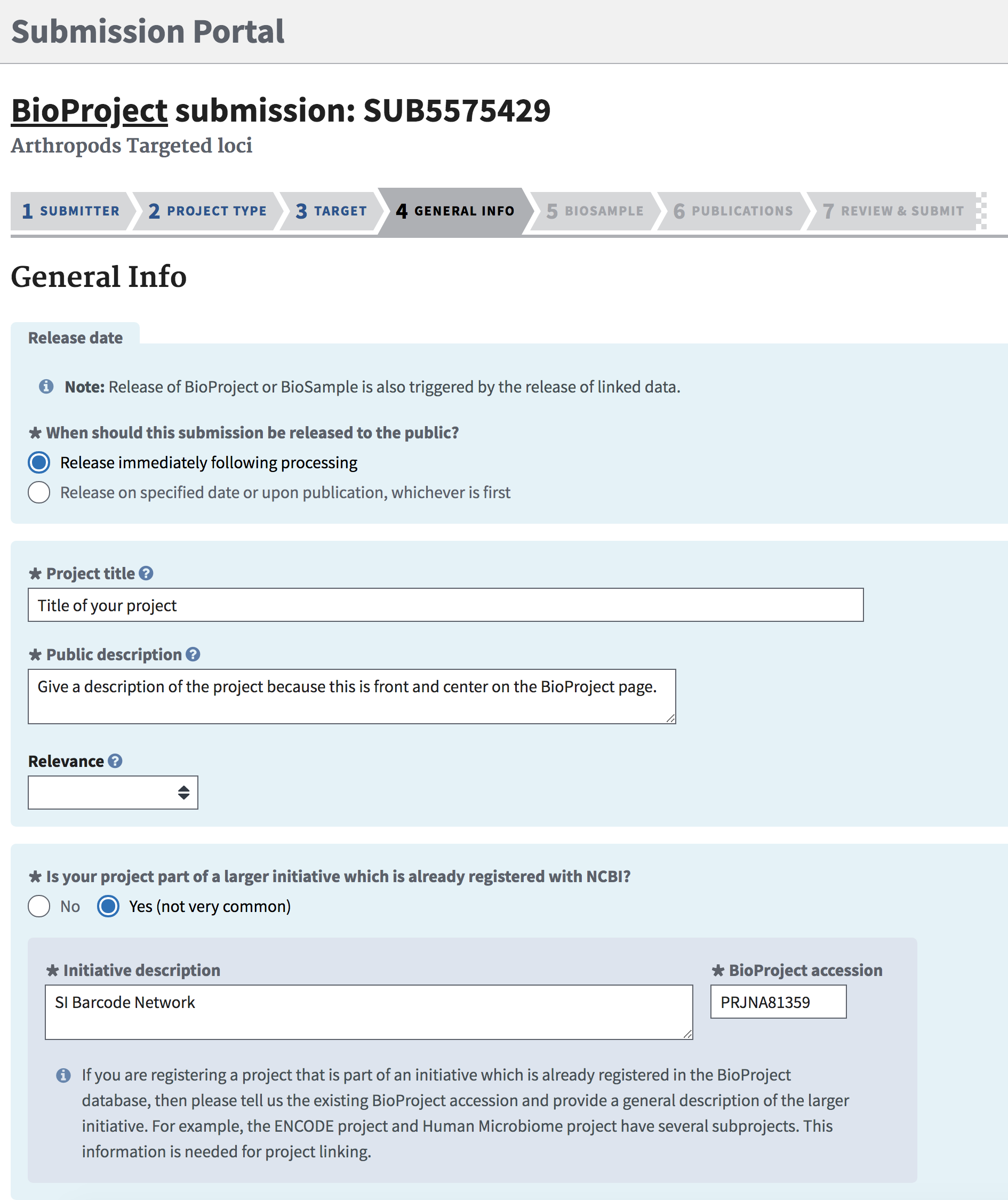
The submission portal will create an automated Project Title based on previous entries, but this can be overwriten with the title of choice for the project.
Give a good description of the project in “Public description”, because this will be front-and-center on the BioProject page.
“Relevance” is not required have a value chosen, but for SIBN funded project this will typically be either “Environmental” or “Evolution”.
Finally, check the “Yes” box to indicate that this project is part of a larger initiative.
If this BioProject falls under the SI Barcode Network, then enter “SI Barcode Network” for Initiative description, and “PRJNA81359” for BioProject Accession.
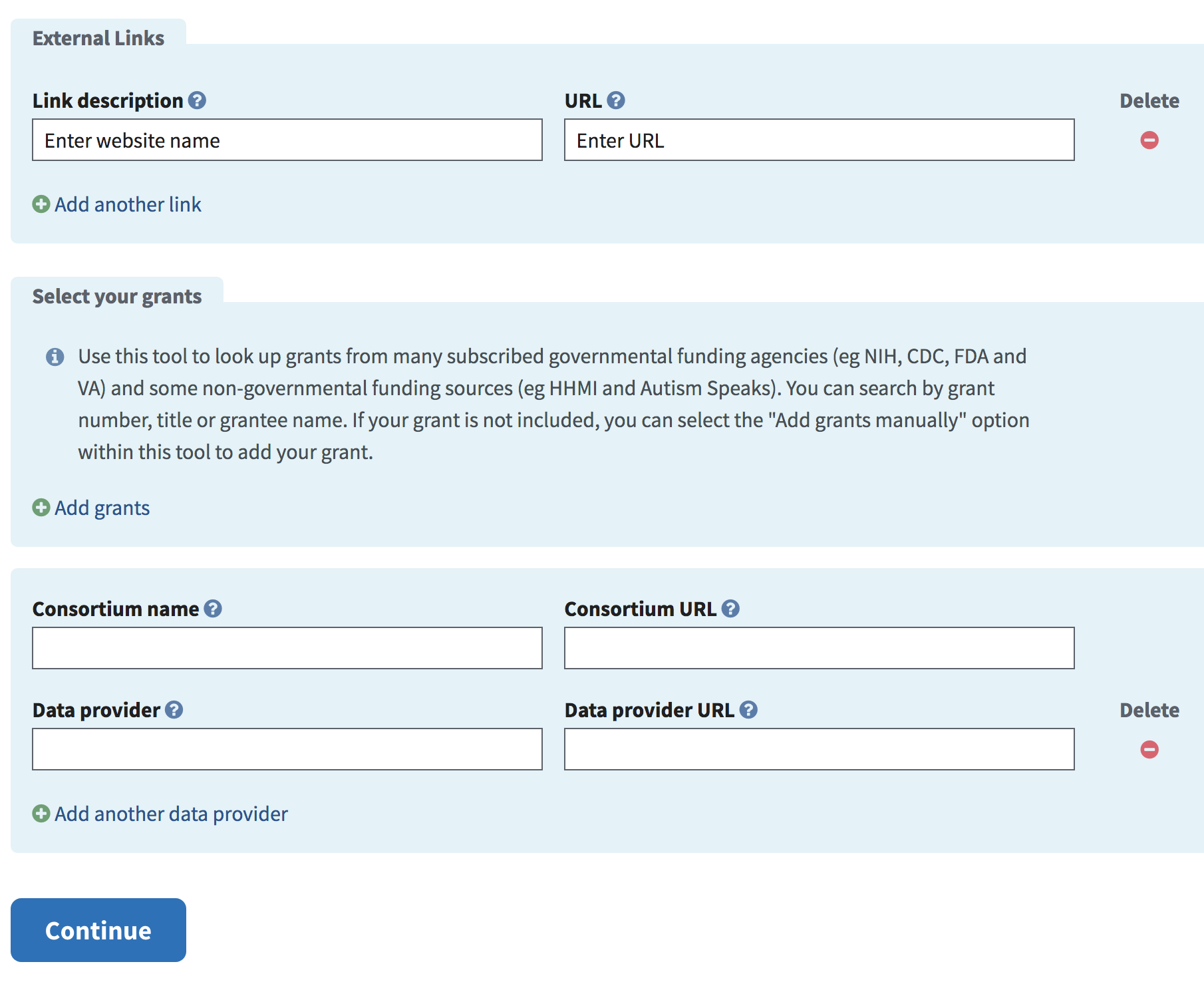
Enter any links to be displayed as part of the BioProject. Add the Consortium and/or Data provider, if applicable.
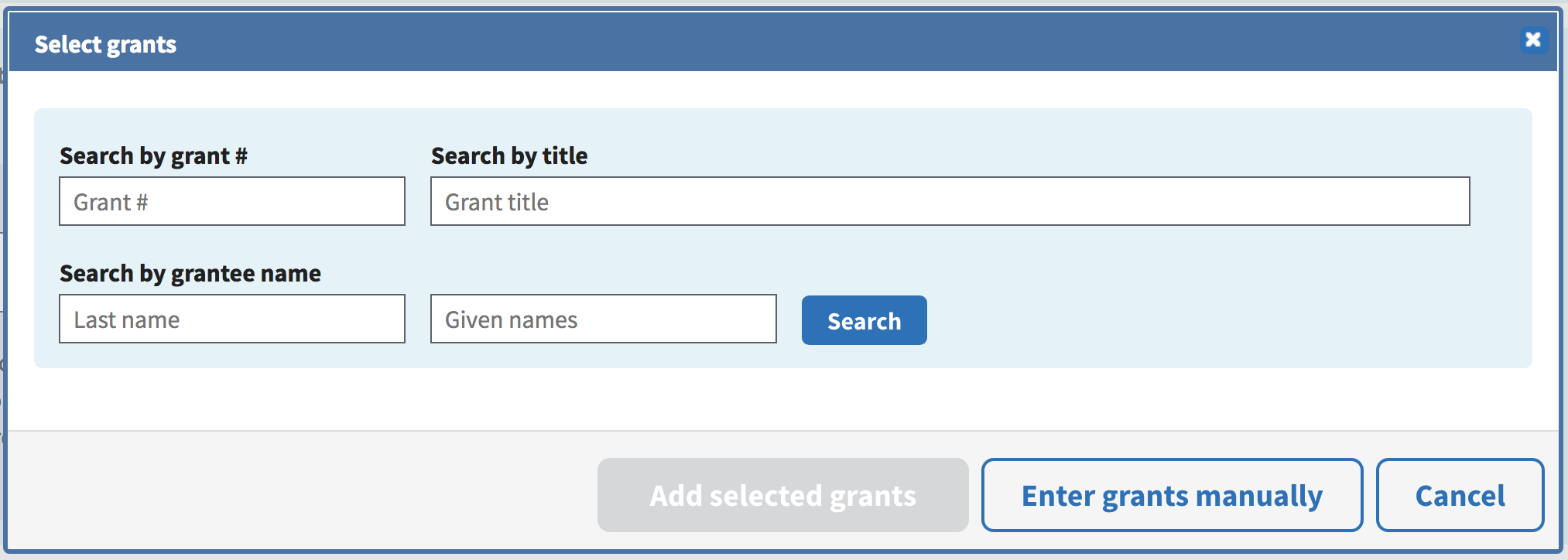
To enter any grants, click the Add grants link to enter the relevant information.
Click “Continue” when ready.
Biosample Tab
SIBN funded projects are not required to create biosamples for sequenced samples, so skip the BioSample page.
Publications Tab
Add any Publications the project has generated. Publications can always be added back in later.
Review & Submit Tab
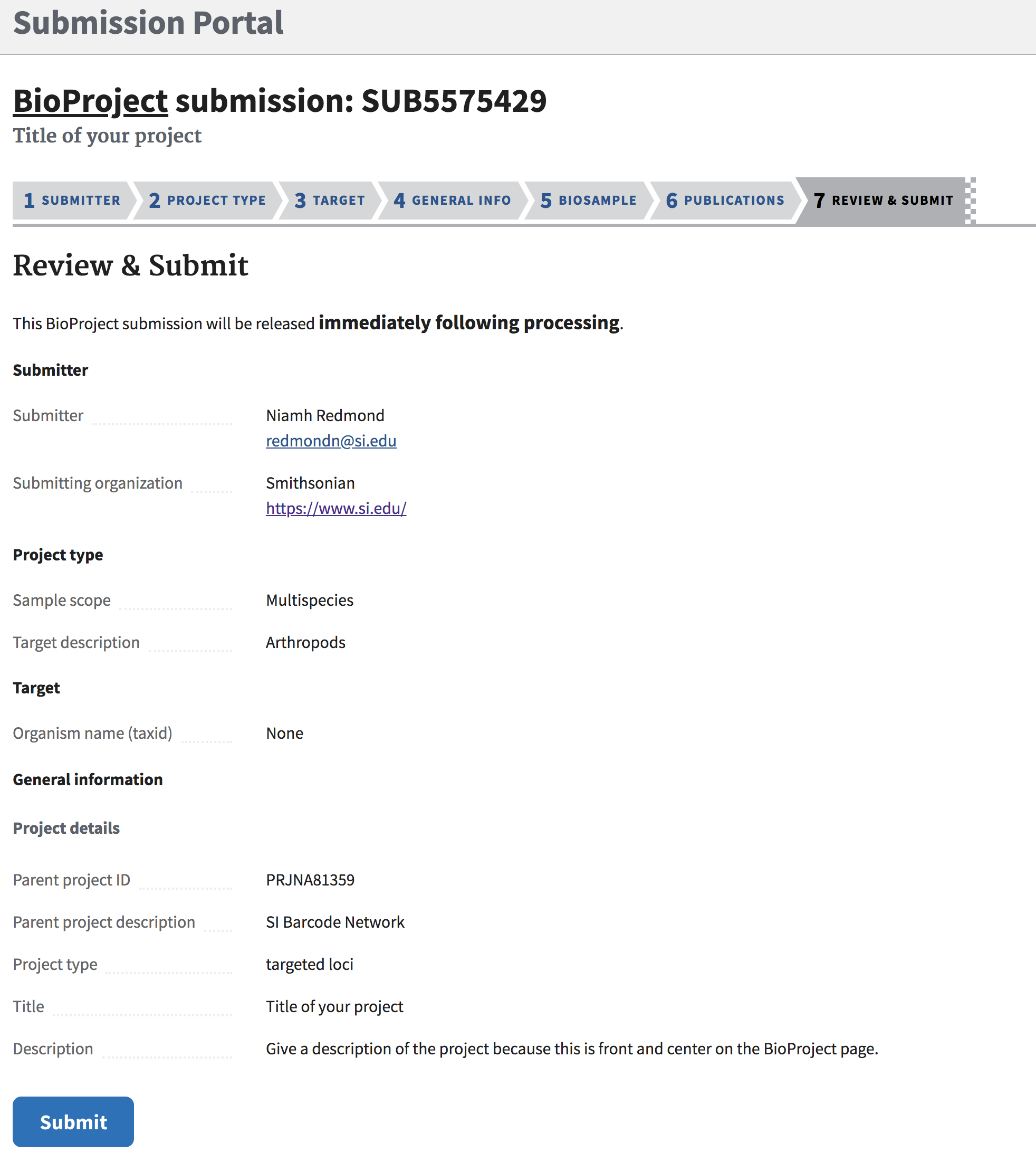
All BioProject data that has been entered is summarized in one place for review. This will be the last chance to make any changes before submitting.
Shortly afterwards, NCBI will send an email to note that the BioProject has been successfully created. Most importantly, they will send the BioProject ID, which can then be added to existing GenBank records or include in new GenBank submissions.
How to Update BioProject Information
If a BioProject has already been published and data need to be updated (i.e any typo corrections or perhaps an addition of a publication), log into the NCBI Submission Portal, navigate to the “My submissions” tab and it should bring up a list of BioProject submissions.
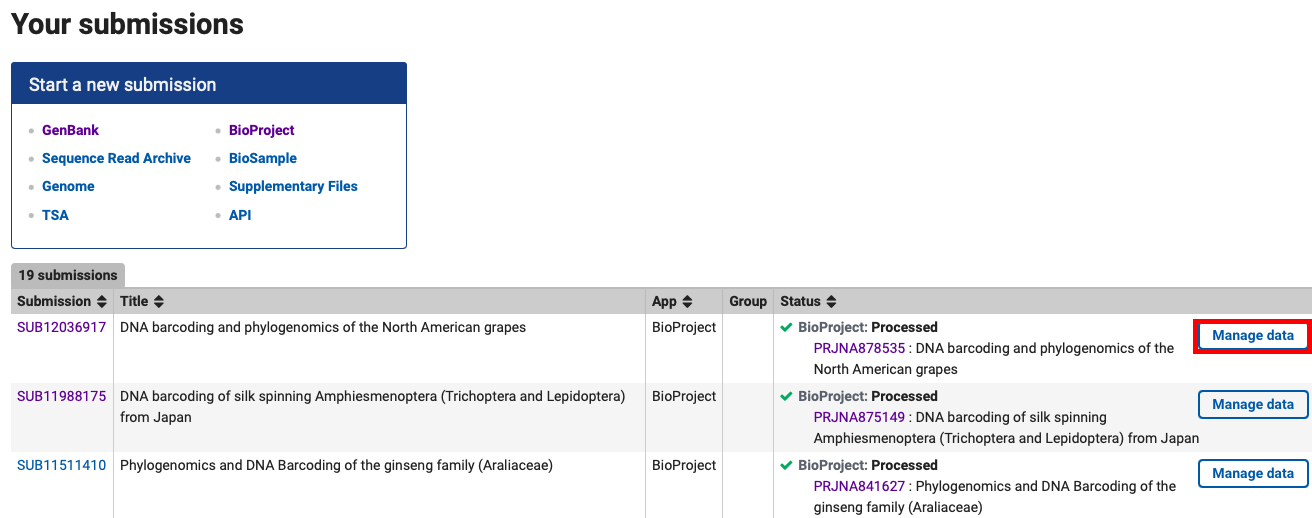

From the list of processed projects, click “Manage Data” to right in the “Status” column. Most changes can be applied by user directly to the BioProject here.
However, if any changes are needed that cannot be made here, email the update request to bioprojecthelp@ncbi.nlm.nih.gov.
Adding a BioProject to Existing GenBank Records
Adding a BioProject ID to sequence records that are already published to GenBank is a manual procedure done through email. There are two options:
Either - Email bioprojecthelp@ncbi.nlm.nih.gov with:
the BioProject ID in the subject line
the range of GenBank accessions to be added to the BioProject in the body of the email
- Or - Treat the BioProject as a source modifier update to the GenBank accessions and email gb-admin@ncbi.nlm.nih.gov with:
the range of GenBank accessions to be updated in the subject line
attach a text file table that contains the fields “acc. num.” and “bioproject” (without the quotations)
Adding a BioProject to New GenBank Submissions
Out of the several different methods of publishing sequences to GenBank (GenBank Submission Portal, BankIt, Sequin, tbl2asn, Geneious, and BOLD), only the Genbank Submission Portal and tbl2asn have methods for adding a BioProject ID to a batch submission.
If submitting metazoan C01 or rDNA through the Genbank Submission Portal, when creating the source modifier table for upload to the portal, simply add a column containing the BioProject ID with the column header “Bioproject” (without the quotations).
If submitting through tbl2asn, follow instructions in the section below for BioProject addition.
For other submission methods, submit the sequences first and treat the sequences as “existing Genbank records” (see above).
tbl2asn
In the tbl2asn instruction manual at https://www.ncbi.nlm.nih.gov/genbank/tbl2asn2/, the 3 files required to create a submission package are a “template file”, a FASTA file containing nucleotide sequences, and a feature table with annotations. The template file is where the BioProject ID is included for a submission.
To create a GenBank submission template file, go to https://submit.ncbi.nlm.nih.gov/genbank/template/submission/, and fill out the form. The last section of the form is for “BioProject/BioSample Information”, and this is where to add the BioProject ID.
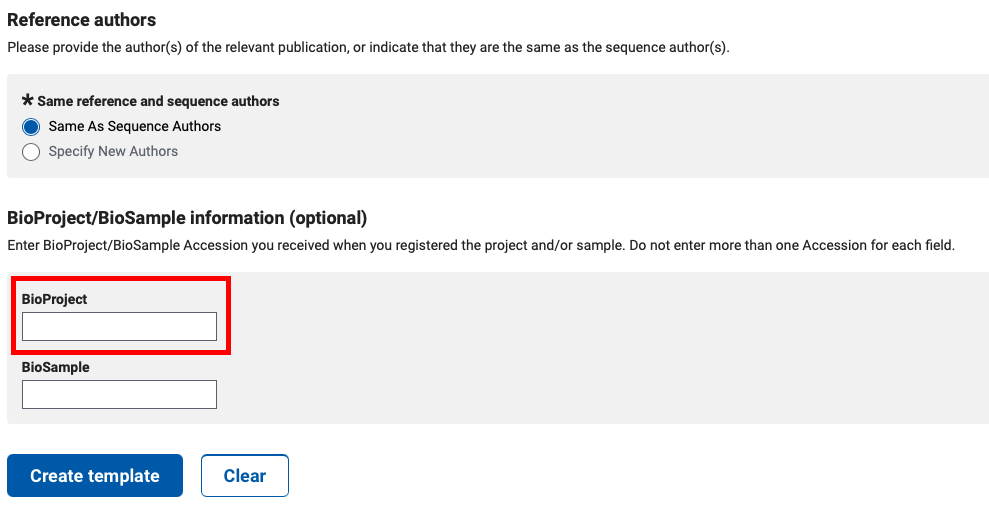
Press the “Create Template” button to download a “.sbt” file, and bundle that with the other components for the tbl2asn command line utility.